Analyze: Selected Oligonucleotides & Sets
|
![]()
Analyze: Selected Oligonucleotides & Sets
|
![]()
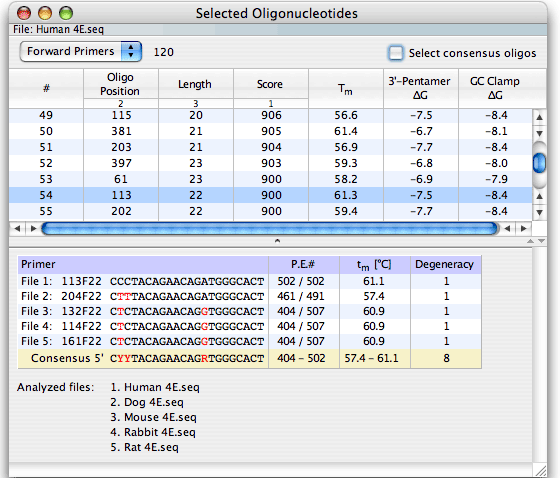
The Analyze - Selected Oligonucleotides window displays a table listing all oligonucleotide selections from the most recent search. Choose the kind of oligos to display (Forward Primers, Upper Probes, etc.) with the button at the upper left corner of the window. At the default setting, the window sorts oligonucleotides by descending score, ascending position and descending length. You may easily change the sort order. The data table lists Tm of the oligo, the 3'-terminal pentamer ΔG (specificity), and GC clamp ΔG stability (the most stable pentamer in the oligo). If you’ve searched for consensus primers or probes, you may pull down the window separator to reveal all sequence alignments and the consensus sequence, as shown above.
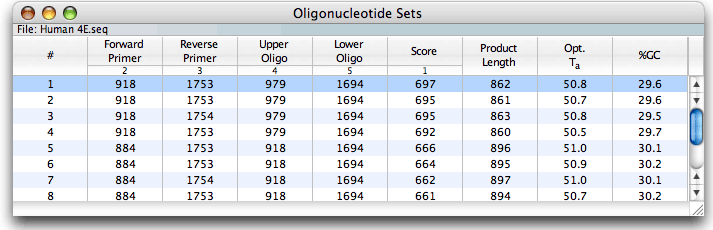
The Analyze - Oligonucleotide Sets window contains a table of primer pairs data from the most recent PCR primer, TaqMan, Molecular Beacons or nested primers search. The following information is included in the table:
• The primer/probe set number
• The primers/probe positions on the + and - strand active sequences
• The PCR product score
• The length of the PCR product the pair would generate
• The calculated optimal annealing temperature recommended for PCR using this primer pair
• The percent GC content of the PCR product generated by this pair
By clicking on any of the listed primer pairs & probes rows in the table, you select the listed sets. Double clicking on a pair also calls up the PCR window that displays a graphic of the PCR product's location on the active sequence, along with other data for running PCR with the selected primer pair. The Sequence window is also updated at the same time.