Analyze: Duplex Formation
|
![]()
Analyze: Duplex Formation
|
![]()

These windows display potential duplexes in a single selected oligo or between various oligonucleotides.
ΔG and hairpin loop Tm values are calculated and displayed in association with each duplex (dimer or hairpin). The stability values of the most stable duplex (shown in red) in each alignment, and of the hairpin structure are expressed in kcal/mol. Tm of hairpins and duplexis are displayed only when they are greater than 0°.
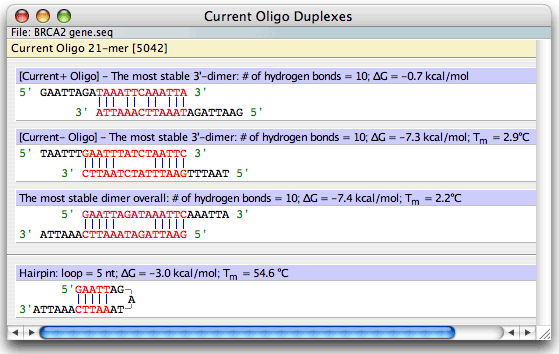
The "Current Oligo" duplex formation window displays:
• The most stable 3' terminal dimer alignment of the current positive strand oligo
• The most stable 3' terminal dimer alignment of the current negative strand oligo
• The most stable dimer alignment overall in the current positive strand oligo
• The most stable hairpin structure in the positive strand of the Current Oligo